DO-MS
Data-Driven Optimization of Mass Spectrometry Methods

Get started now Download JPR Articles GitHub Repository
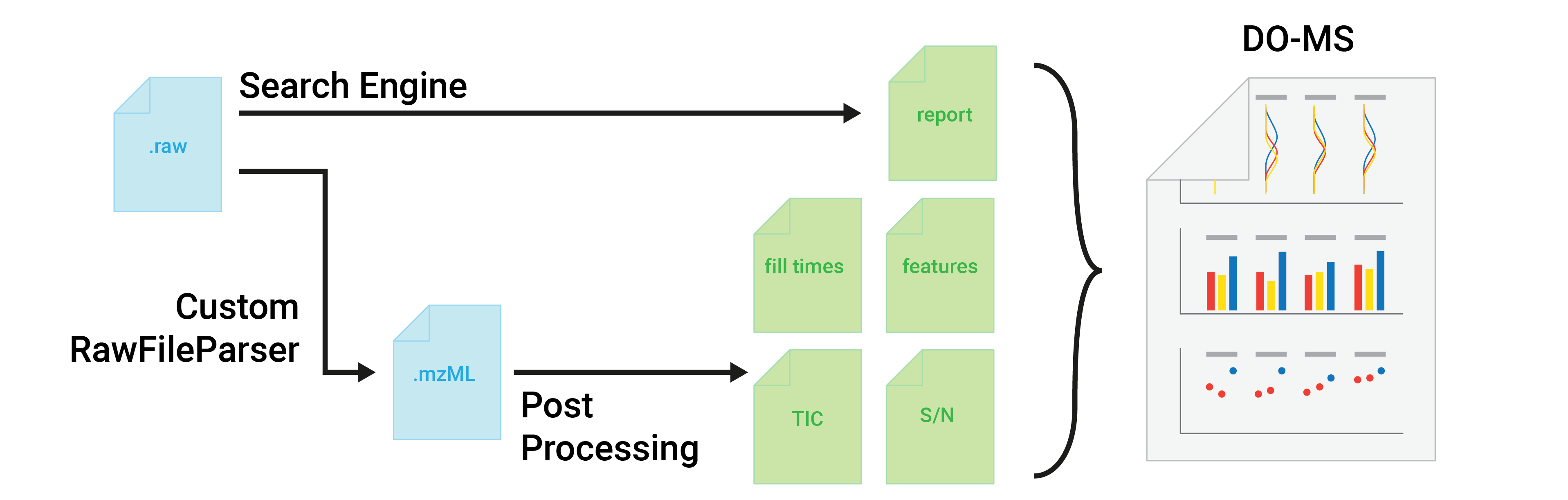
Aim of DO-MS
The performance of ultrasensitive liquid chromatography and tandem mass spectrometry (LC-MS/MS) methods, such as single-cell proteomics by mass spectrometry and multiplexed data-independent acquisition experiments, depends on multiple interdependent parameters. This interdependence makes it challenging to specifically pinpoint the sources of problems in the LC-MS/MS methods.
This applies to data-dependent acquisition (DDA) as well as to data-indepent acquisition (DIA) experiments. For example, a low signal at the MS2 level in a DDA experiment can be due to poor LC separation, ionization, apex targeting, ion transfer, or ion detection. DO-MS aims to specifically diagnose such problems by interactively visualizing data from all levels of bottom-up LC-MS/MS analysis.
Research articles
The development and applications of DO-MS were described in two research articles published in the Journal of Proteome Research. The first version from 2019 focussed on DDA data and the second version from 2023 focussed on DIA data.
-
Wallmann G., Leduc A., Slavov N. Data-Driven Optimization of DIA Mass-Spectrometry by DO-MS, J. Proteome Res. doi: 10.1021/acs.jproteome.3c00177 (2023) preprint PDF -
Huffman RG, Specht H, Chen AT, Slavov N. DO-MS: Data-Driven Optimization of Mass Spectrometry Methods J. of Proteome Res. doi: 10.1021/acs.jproteome.9b00039 (2019) preprint PDF
Installation
Install this application by downloading it from the release page and by following the installation instructions.
Getting Started
Please read our detailed getting started guides:
- Getting started with DIA preprocessing
- Getting started with DIA reports
- Getting started with DDA reports
Requirements
This application has been tested on R >= 3.5.0, OSX 10.14 / Windows 7/8/10/11. R can be downloaded from the main R Project page or downloaded with the RStudio Application. All modules are maintained for MaxQuant >= 1.6.0.16 and DIA-NN > 1.8.1.
The application suffers from visual glitches when displayed on unsupported older browsers (such as IE9 commonly packaged with RStudio on Windows). Please use IE >= 11, Firefox, or Chrome for the best user experience.
Running the Interactive Application
The easiest way to run the app is directly through RStudio, by opening the DO-MS.Rproj Rproject file
and clicking the “Run App” button at the top of the application, after opening the server.R file. We recommend checking the “Run External” option to open the application in your default browser instead of the RStudio Viewer.
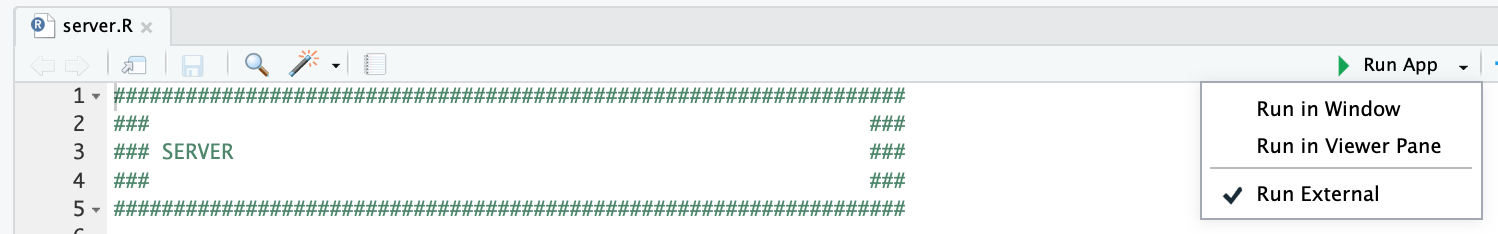
You can also start the application by running the start_server.R script.
Customization
DO-MS is designed to be easily user-customizable for in-house proteomics workflows. Please see Building Your Own Modules for more details.
Hosting as a Server
Please see Hosting as a Server for more details.
Supporting other Search Engines
This application is currently maintained for MaxQuant >= 1.6.0.16 and DIA-NN >= 1.8. Adapting to other search engines is possible but not provided out-of-the-box. Please see Integrating Other Search Engines for more details.
Can I use this for Metabolomics, Lipidomics, etc… ?
While the base library of modules are based around bottom-up proteomics by LC-MS/MS, this project is fundamentally compatible with any delimited text files (CSV, TSV, etc). These implementations will require some programming work, but once it is done DO-MS gives you a extensible framework that can be used over-and-over again to generate shareable reports. See Integrating Other Search Engines for more details
About the project
The manuscripts for this tool is published at the Journal of Proteome Research and bioRxiv. The research has been supported by funding from the NIH Director’s Award by an Allen Distinguished Investigator Award from the Paul G. Allen Frontiers Group.
Contact the authors by email: nslavov{at}northeastern.edu.
License
DO-MS is distributed by an MIT license.
Contributing
Please feel free to contribute to this project by opening an issue or pull request in the GitHub repository.
Data Availability
DO-MS reports and example data can be found Here. All raw data and search engine results from the DO-MS DIA paper are avilable on MassIVE under the following id: MSV000091733.
Help!
For any bugs, questions, or feature requests, please use the GitHub issue system to contact the developers.
